In this document, we take great care providing all steps and R codes required to check whether our matching procedure allowed to improve covariates balance. We compare days where:
- treated units are days where winds blow from the North-East in t.
- control units are day winds blow from other directions in t.
We adjust for calendar indicators and weather confouding factors.
Should you have any questions, need help to reproduce the analysis or find coding errors, please do not hesitate to contact me at leo.zabrocki@psemail.eu
Required Packages
To reproduce exactly the 4_script_checking_balance_improvement.html document, we first need to have installed:
- the R programming language
- RStudio, an integrated development environment for R, which will allow you to knit the
4_script_checking_balance_improvement.Rmdfile and interact with the R code chunks - the R Markdown package
- and the Distill package which provides the template for this document.
Once everything is set up, we have to load the following packages:
# load required packages
library(knitr) # for creating the R Markdown document
library(here) # for files paths organization
library(tidyverse) # for data manipulation and visualization
library(ggridges) # for ridge density plots
library(Cairo) # for printing custom police of graphs
library(patchwork) # combining plots
We finally load our custom ggplot2 theme for graphs:
Preparing the Data
We load the initial and matched data and bind them together:
# load matching data
data_matching <-
readRDS(here::here("inputs", "1.data", "5.matched_data", "matching_data.rds")) %>%
mutate(dataset = "Initial Data")
# load matched data
data_matched <-
readRDS(here::here("inputs", "1.data", "5.matched_data", "matched_data.rds")) %>%
mutate(dataset = "Matched Data")
# bind the two datasets
data <- bind_rows(data_matching, data_matched)
We change labels of the is_treated variable :
data <- data %>%
mutate(is_treated = ifelse(is_treated == "TRUE", "True", "False"))
Love Plots
Continuous Weather Covariates
Please show me the code!
# compute figures for the love plot
data_weather_continuous <- data %>%
select(
dataset,
is_treated,
contains("temperature"),
contains("humidity"),
contains("wind_speed")
) %>%
pivot_longer(
cols = -c(is_treated, dataset),
names_to = "variable",
values_to = "values"
) %>%
mutate(
weather_variable = NA %>%
ifelse(
str_detect(variable, "temperature_average"),
"Average Temperature",
.
) %>%
ifelse(
str_detect(variable, "humidity_average"),
"Humidity Average",
.
) %>%
ifelse(str_detect(variable, "wind_speed"), "Wind Speed", .)
) %>%
mutate(time = "0" %>%
ifelse(str_detect(variable, "lag_1"), "-1", .) %>%
ifelse(str_detect(variable, "lead_1"), "+1", .)) %>%
filter(time != "+1") %>%
mutate(time = fct_relevel(time, "-1", "0")) %>%
select(dataset, is_treated, weather_variable, time, values)
data_abs_difference_continuous_weather <-
data_weather_continuous %>%
group_by(dataset, weather_variable, time, is_treated) %>%
summarise(mean_values = mean(values, na.rm = TRUE)) %>%
summarise(abs_difference = abs(mean_values[2] - mean_values[1]))
data_sd_weather_continuous <- data_weather_continuous %>%
filter(dataset== "Initial Data" & is_treated == "True") %>%
group_by(dataset, weather_variable, time, is_treated) %>%
summarise(sd_treatment = sd(values, na.rm = TRUE)) %>%
ungroup() %>%
select(weather_variable, time, sd_treatment)
data_love_continuous_weather <-
left_join(
data_abs_difference_continuous_weather,
data_sd_weather_continuous,
by = c("weather_variable", "time")
) %>%
mutate(standardized_difference = abs_difference / sd_treatment) %>%
select(-c(abs_difference, sd_treatment))
# make the graph
graph_love_plot_continuous_weather <-
ggplot(
data_love_continuous_weather,
aes(
y = time,
x = standardized_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)
) +
geom_vline(xintercept = 0, size = 0.3) +
geom_vline(xintercept = 0.1,
color = "black",
linetype = "dashed") +
geom_point(size = 4, alpha = 0.8) +
scale_x_continuous(breaks = scales::pretty_breaks(n = 5)) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
facet_wrap( ~ weather_variable, scales = "free_y") +
xlab("Standardized Mean Differences") +
ylab("Day") +
theme_tufte()
# plot the graph
graph_love_plot_continuous_weather
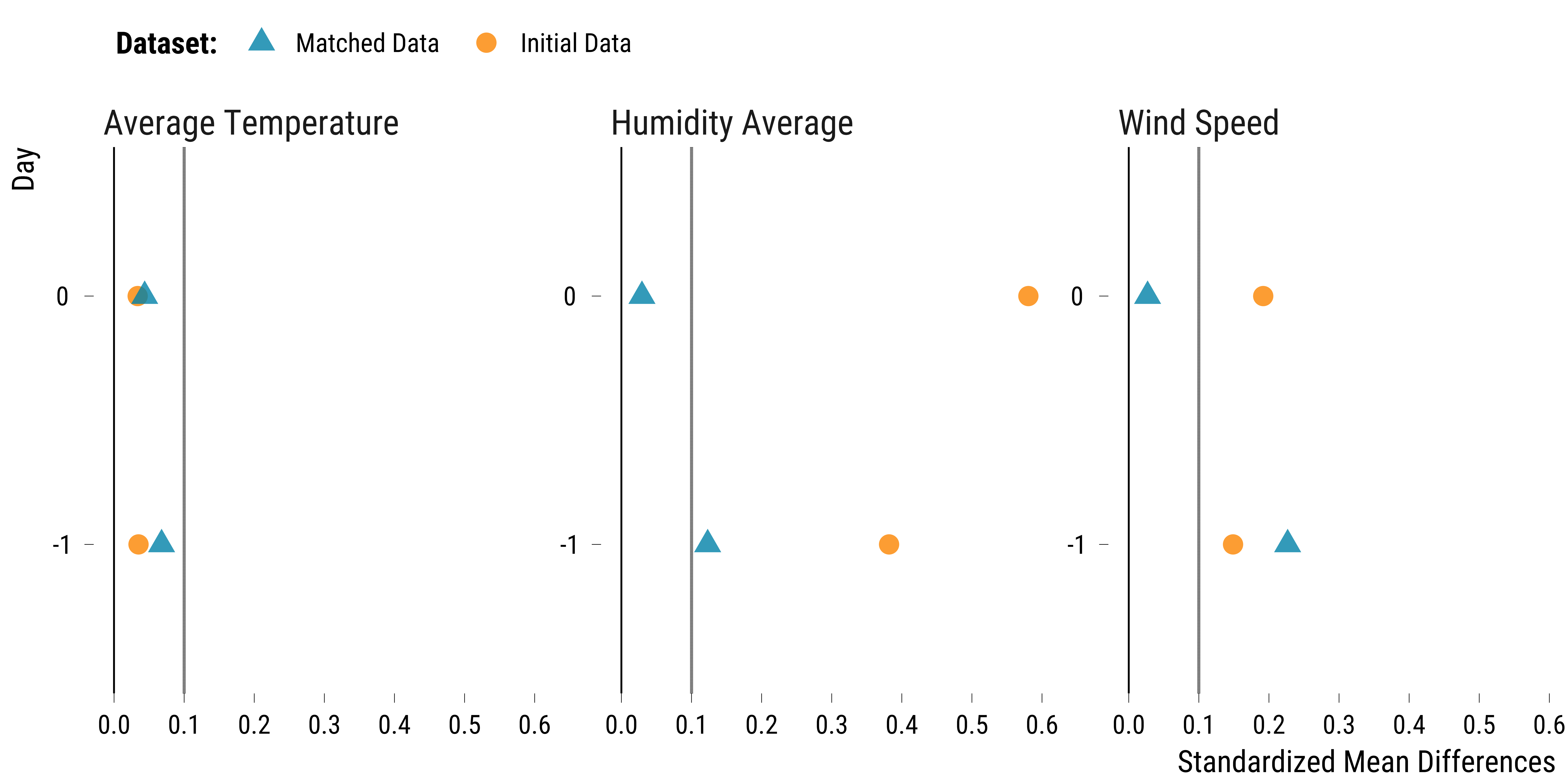
Please show me the code!
# save the graph
ggsave(
graph_love_plot_continuous_weather,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_continuous_weather.pdf"
),
width = 30,
height = 12,
units = "cm",
device = cairo_pdf
)
Categorical Weather Covariates
Please show me the code!
# compute figures for the love plot
data_weather_categorical <- data %>%
select(
dataset,
is_treated,
contains("rainfall_duration"),
"wind_direction_categories_lag_1"
) %>%
drop_na() %>%
mutate_all( ~ as.character(.)) %>%
pivot_longer(
cols = -c(dataset, is_treated),
names_to = "variable",
values_to = "values"
) %>%
# group by is_treated, variable and values
group_by(dataset, is_treated, variable, values) %>%
# compute the number of observations
summarise(n = n()) %>%
# compute the proportion
mutate(freq = round(n / sum(n) * 100, 0)) %>%
ungroup() %>%
mutate(
weather_variable = NA %>%
ifelse(str_detect(variable, "wind"), "Wind Direction", .) %>%
ifelse(str_detect(variable, "rainfall"), "Rainfall Duration", .)
) %>%
mutate(time = "t" %>%
ifelse(str_detect(variable, "lag_1"), "t-1", .) %>%
ifelse(str_detect(variable, "lead_1"), "t+1", .)) %>%
filter(time != "t+1") %>%
mutate(variable = paste(weather_variable, time, sep = " ")) %>%
select(dataset, is_treated, weather_variable, variable, values, freq) %>%
pivot_wider(names_from = is_treated, values_from = freq) %>%
mutate(abs_difference = abs(`True` - `False`)) %>%
filter(weather_variable != "Wind Direction t")
# create the figure for wind direction
graph_love_plot_wind_direction <- data_weather_categorical %>%
filter(weather_variable == "Wind Direction") %>%
ggplot(.,
aes(
y = fct_rev(values),
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
facet_wrap( ~ variable, scales = "free_y", ncol = 3) +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")))
# print the figure for wind direction
graph_love_plot_wind_direction

Please show me the code!
# save the figure for wind direction
ggsave(
graph_love_plot_wind_direction,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_wind_direction.pdf"
),
width = 20,
height = 10,
units = "cm",
device = cairo_pdf
)
# create the figure for rainfall dummy
graph_love_plot_rainfall <- data_weather_categorical %>%
filter(weather_variable == "Rainfall Duration") %>%
mutate(variable = fct_relevel(variable, "Rainfall Duration t-1", "Rainfall Duration t")) %>%
ggplot(.,
aes(
y = fct_rev(values),
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
facet_wrap( ~ variable) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")),
axis.text.y = element_text(hjust = 1))
# print the figure for rainfall dummy
graph_love_plot_rainfall
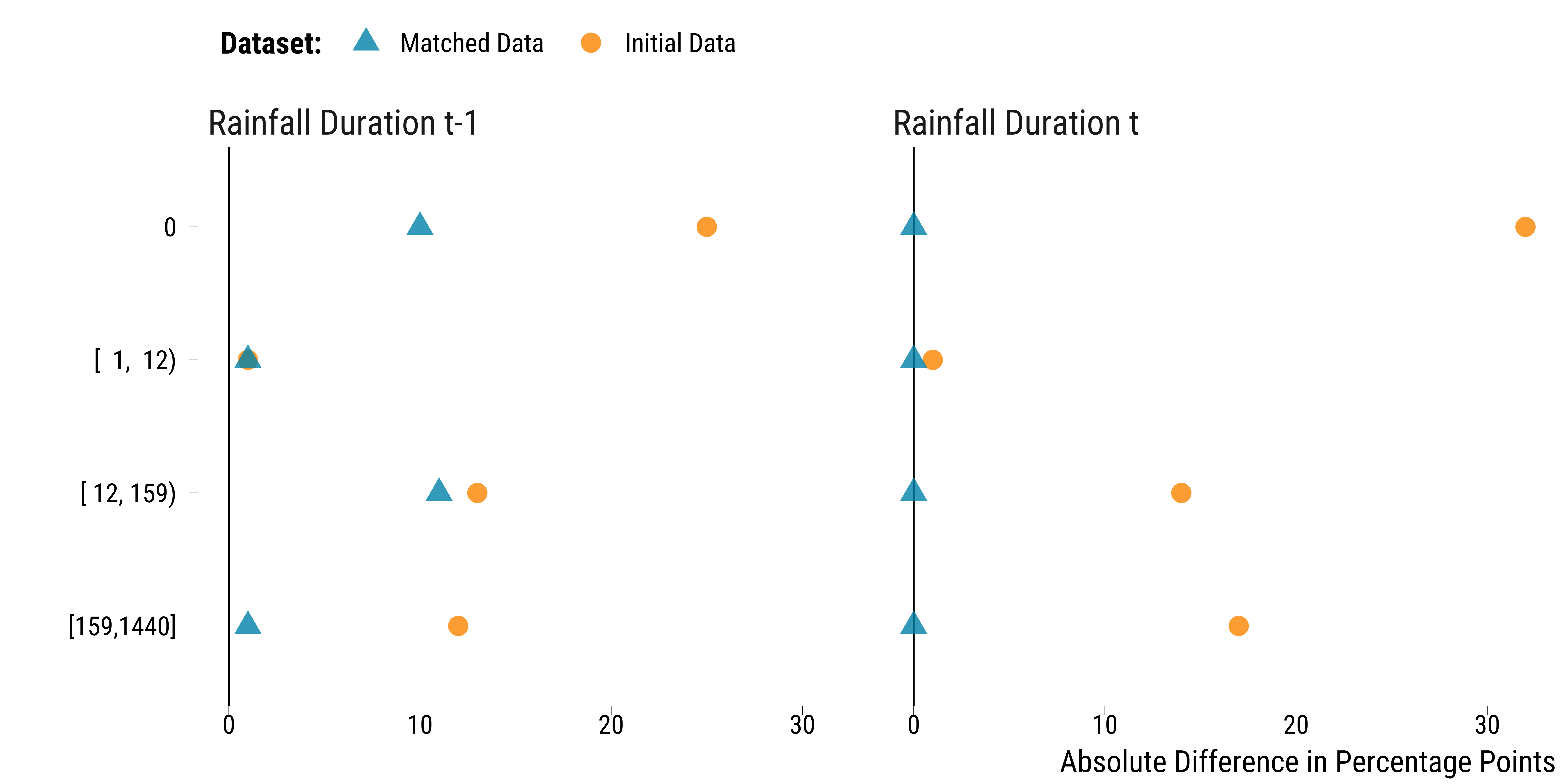
Please show me the code!
# save the figure for rainfall dummy
ggsave(
graph_love_plot_rainfall,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_rainfall.pdf"
),
width = 20,
height = 10,
units = "cm",
device = cairo_pdf
)
Pollutants
Please show me the code!
# compute figures for the love plot
data_pollutants <- data %>%
select(
dataset,
is_treated,
mean_no2_lag_1,
mean_o3_lag_1,
mean_pm10_lag_1,
mean_pm25_lag_1
) %>%
pivot_longer(
cols = -c(dataset, is_treated),
names_to = "variable",
values_to = "values"
) %>%
mutate(
pollutant = NA %>%
ifelse(str_detect(variable, "no2"), "NO2 in t-1", .) %>%
ifelse(str_detect(variable, "o3"), "O3 in t-1", .) %>%
ifelse(str_detect(variable, "pm10"), "PM10 in t-1", .) %>%
ifelse(str_detect(variable, "pm25"), "PM2.5 in t-1", .)
) %>%
select(dataset, is_treated, pollutant, values)
data_abs_difference_pollutants <- data_pollutants %>%
group_by(dataset, pollutant, is_treated) %>%
summarise(mean_values = mean(values, na.rm = TRUE)) %>%
summarise(abs_difference = abs(mean_values[2] - mean_values[1]))
data_sd_pollutants <- data_pollutants %>%
filter(dataset== "Initial Data" & is_treated == "True") %>%
group_by(pollutant, is_treated) %>%
summarise(sd_treatment = sd(values, na.rm = TRUE)) %>%
ungroup() %>%
select(pollutant, sd_treatment)
data_love_pollutants <-
left_join(data_abs_difference_pollutants,
data_sd_pollutants,
by = c("pollutant")) %>%
mutate(standardized_difference = abs_difference / sd_treatment) %>%
select(-c(abs_difference, sd_treatment))
# create the graph
graph_love_plot_pollutants <-
ggplot(
data_love_pollutants,
aes(
y = fct_rev(pollutant),
x = standardized_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)
) +
geom_vline(xintercept = 0, size = 0.3) +
geom_vline(xintercept = 0.1,
color = "black",
linetype = "dashed") +
geom_point(size = 4, alpha = 0.8) +
scale_x_continuous(breaks = scales::pretty_breaks(n = 5)) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
xlab("Standardized Mean Differences") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")),
axis.text.y = element_text(hjust = 1))
# print the graph
graph_love_plot_pollutants
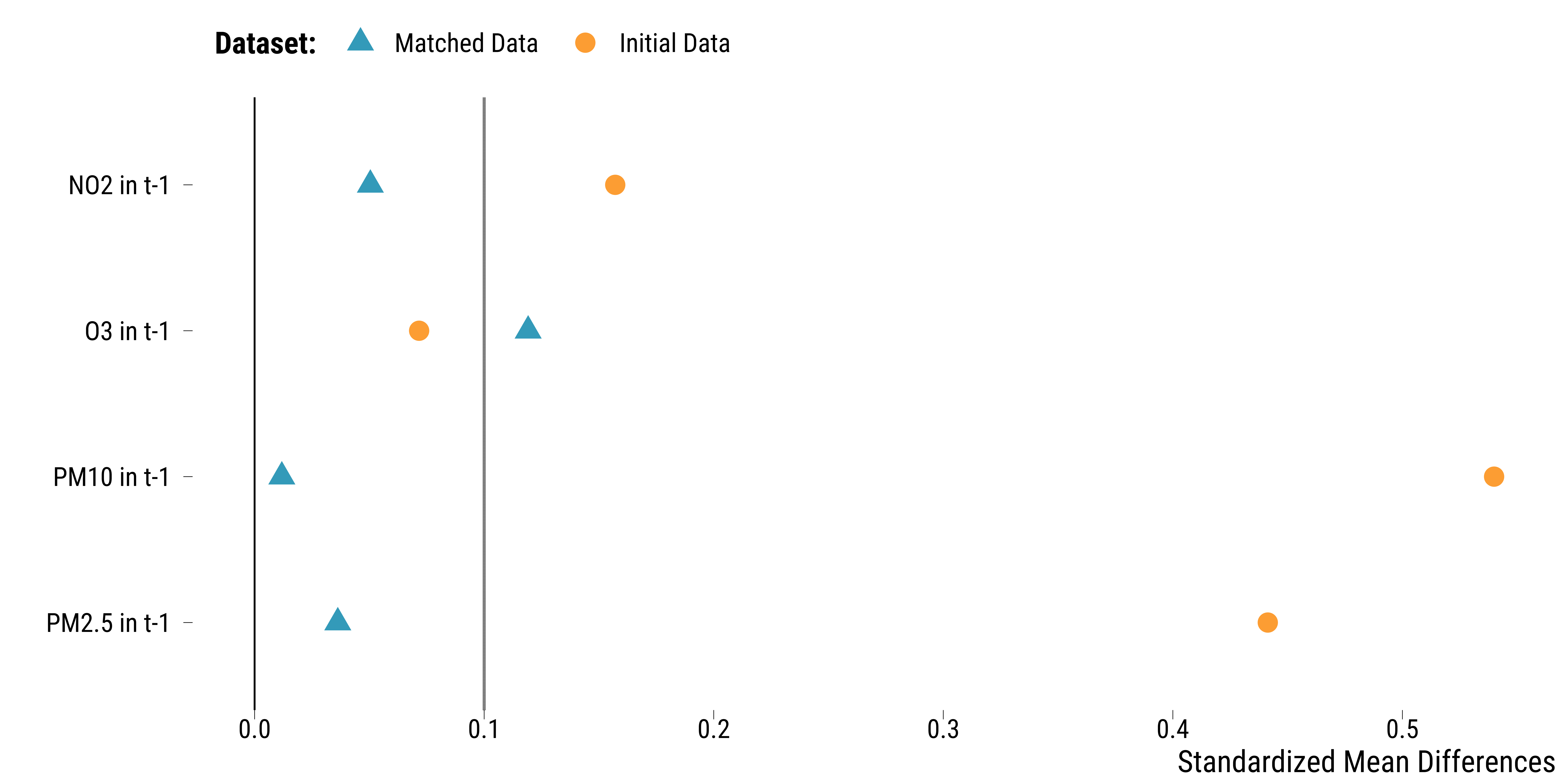
Please show me the code!
# save the graph
ggsave(
graph_love_plot_pollutants,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_pollutants.pdf"
),
width = 20,
height = 10,
units = "cm",
device = cairo_pdf
)
Calendar Indicators
Create the relevant data:
Please show me the code!
# compute figures for the love plot
data_calendar <- data %>%
mutate(weekday = lubridate::wday(date, abbr = FALSE, label = TRUE)) %>%
select(dataset,
is_treated,
weekday,
holidays_dummy,
bank_day_dummy,
month,
year) %>%
mutate_at(vars(holidays_dummy, bank_day_dummy),
~ ifelse(. == 1, "True", "False")) %>%
mutate_all( ~ as.character(.)) %>%
pivot_longer(
cols = -c(dataset, is_treated),
names_to = "variable",
values_to = "values"
) %>%
# group by is_treated, variable and values
group_by(dataset, is_treated, variable, values) %>%
# compute the number of observations
summarise(n = n()) %>%
# compute the proportion
mutate(freq = round(n / sum(n) * 100, 0)) %>%
ungroup() %>%
mutate(
calendar_variable = NA %>%
ifelse(str_detect(variable, "weekday"), "Day of the Week", .) %>%
ifelse(str_detect(variable, "holidays_dummy"), "Holidays", .) %>%
ifelse(str_detect(variable, "bank_day_dummy"), "Bank Day", .) %>%
ifelse(str_detect(variable, "month"), "Month", .) %>%
ifelse(str_detect(variable, "year"), "Year", .)
) %>%
select(dataset, is_treated, calendar_variable, values, freq) %>%
pivot_wider(names_from = is_treated, values_from = freq) %>%
mutate(abs_difference = abs(`True` - `False`)) %>%
filter(values != "False")
Plot for bank days and holidays:
Please show me the code!
# graph for bank days and holidays
graph_love_plot_bank_holidays <- data_calendar %>%
filter(calendar_variable %in% c("Bank Day", "Holidays")) %>%
ggplot(.,
aes(
y = values,
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
facet_wrap(~ calendar_variable) +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")))
# print the plot
graph_love_plot_bank_holidays
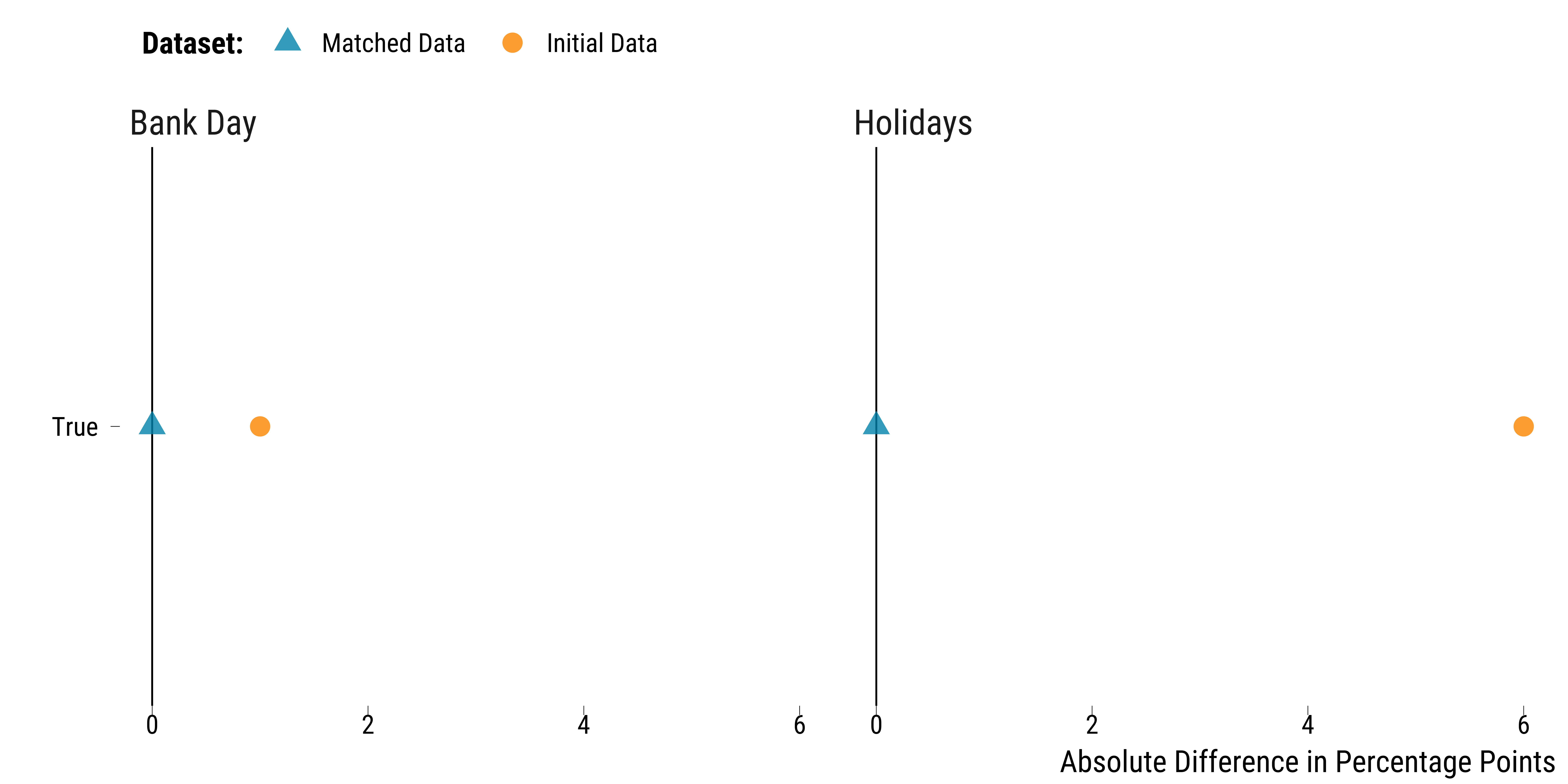
Please show me the code!
# save the plot
ggsave(
graph_love_plot_bank_holidays,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_bank_holidays.pdf"
),
width = 16,
height = 8,
units = "cm",
device = cairo_pdf
)
Plot for days of the week:
Please show me the code!
# graph for weekdays
graph_love_plot_weekday <- data_calendar %>%
filter(calendar_variable == "Day of the Week") %>%
mutate(
values = fct_relevel(
values,
"Monday",
"Tuesday",
"Wednesday",
"Thursday",
"Friday",
"Saturday",
"Sunday"
)
) %>%
ggplot(.,
aes(
y = fct_rev(values),
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")),
axis.text.y = element_text(hjust = 1))
# print the plot
graph_love_plot_weekday

Please show me the code!
# save the plot
ggsave(
graph_love_plot_weekday,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_weekday.pdf"
),
width = 16,
height = 8,
units = "cm",
device = cairo_pdf
)
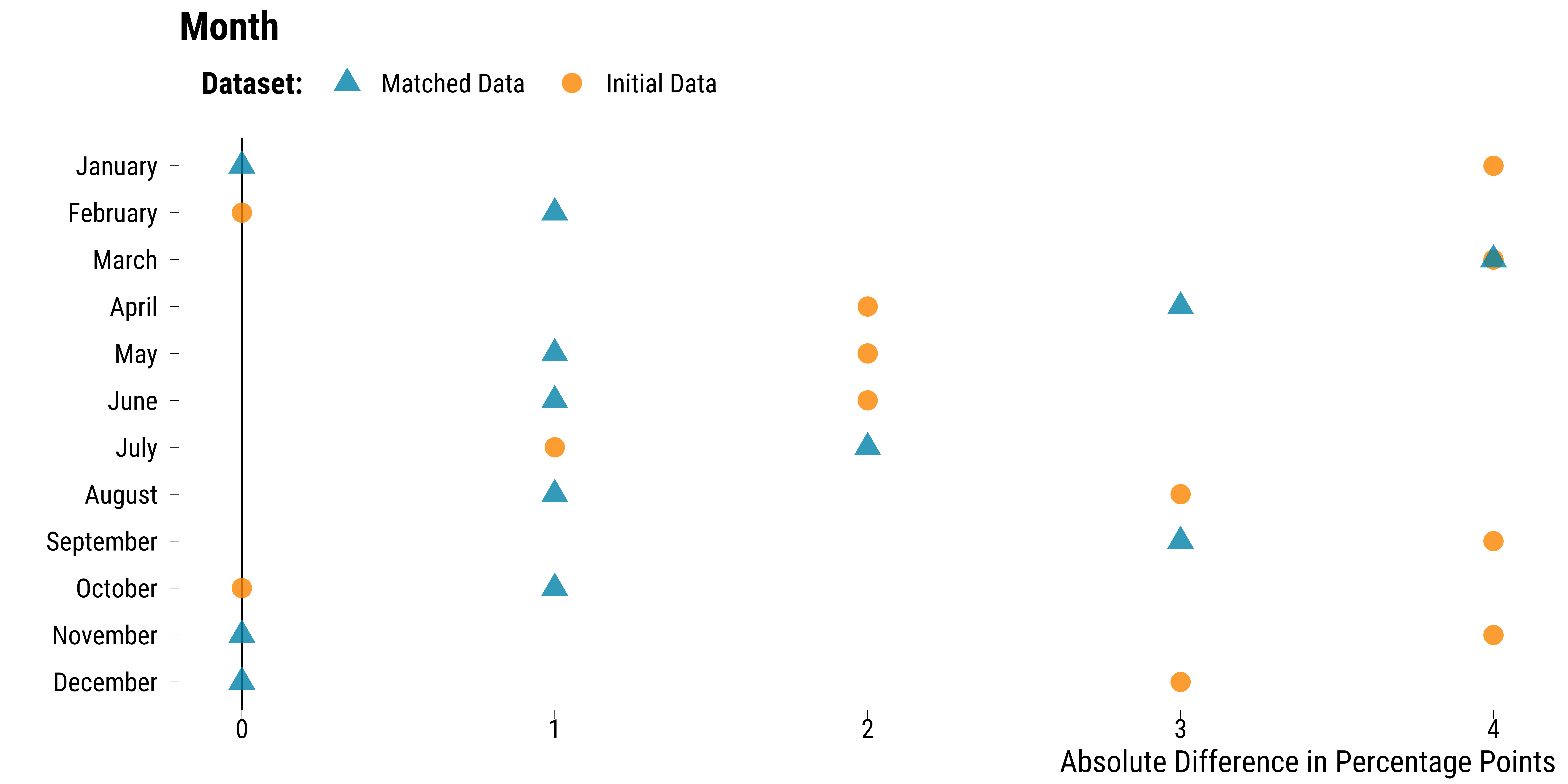
Plot for months:
Please show me the code!
# graph for month
graph_love_plot_month <- data_calendar %>%
filter(calendar_variable == "Month") %>%
mutate(
values = fct_relevel(
values,
"January",
"February",
"March",
"April",
"May",
"June",
"July",
"August",
"September",
"October",
"November",
"December"
)
) %>%
ggplot(.,
aes(
y = fct_rev(values),
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
ggtitle("Month") +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")),
axis.text.y = element_text(hjust = 1))
# print the plot
graph_love_plot_month
Please show me the code!
# save the plot
ggsave(
graph_love_plot_month,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_month.pdf"
),
width = 16,
height = 8,
units = "cm",
device = cairo_pdf
)
Plot for years:
Please show me the code!
# graph for year
graph_love_plot_year <- data_calendar %>%
filter(calendar_variable == "Year") %>%
ggplot(.,
aes(
y = as.factor(as.numeric(values)),
x = abs_difference,
colour = fct_rev(dataset),
shape = fct_rev(dataset)
)) +
geom_vline(xintercept = 0, size = 0.3) +
geom_point(size = 4, alpha = 0.8) +
scale_colour_manual(name = "Dataset:", values = c(my_blue, my_orange)) +
scale_shape_manual(name = "Dataset:", values = c(17, 16)) +
ggtitle("Year") +
xlab("Absolute Difference in Percentage Points") +
ylab("") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")),
axis.text.y = element_text(hjust = 1))
# print the graph
graph_love_plot_year

Please show me the code!
# save the plot
ggsave(
graph_love_plot_year,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_love_plot_year.pdf"
),
width = 16,
height = 8,
units = "cm",
device = cairo_pdf
)
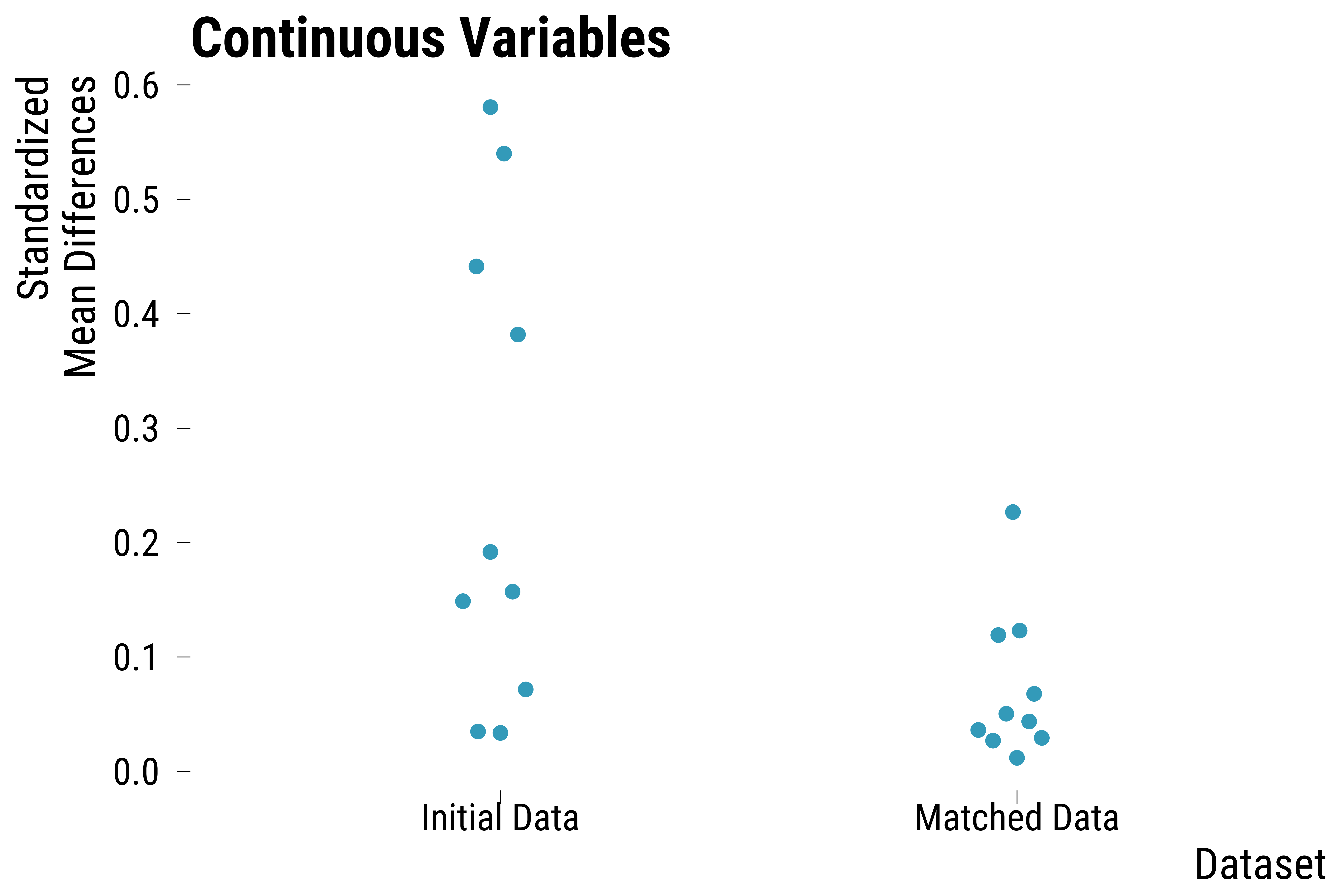
Overall Balance Improvement
We finally plot the distribution of standardized mean differences for continuous covariates or the absolute percentage points differences for categorical covariates between treated and control units before and after matching.
Continuous Covariates
Please show me the code!
# we select the dataset indicator and the standardized difference
data_love_pollutants <- data_love_pollutants %>%
ungroup() %>%
select(dataset, standardized_difference)
data_love_continuous_weather <- data_love_continuous_weather %>%
ungroup() %>%
select(dataset, standardized_difference)
data_continuous_love <-
bind_rows(data_love_pollutants, data_love_continuous_weather)
# create the graph
graph_boxplot_continuous_balance_improvement <-
ggplot(data_continuous_love,
aes(x = dataset, y = standardized_difference)) +
ggbeeswarm::geom_quasirandom(
shape = 16,
size = 2,
width = 0.1,
color = my_blue,
alpha = 0.8
) +
scale_y_continuous(breaks = scales::pretty_breaks(n = 5)) +
xlab("Dataset") +
ylab("Standardized\nMean Differences") +
ggtitle("Continuous Variables") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")))
# print the graph
graph_boxplot_continuous_balance_improvement
Please show me the code!
# save the graph
ggsave(
graph_boxplot_continuous_balance_improvement,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_boxplot_continuous_balance_improvement.pdf"
),
width = 12,
height = 8,
units = "cm",
device = cairo_pdf
)
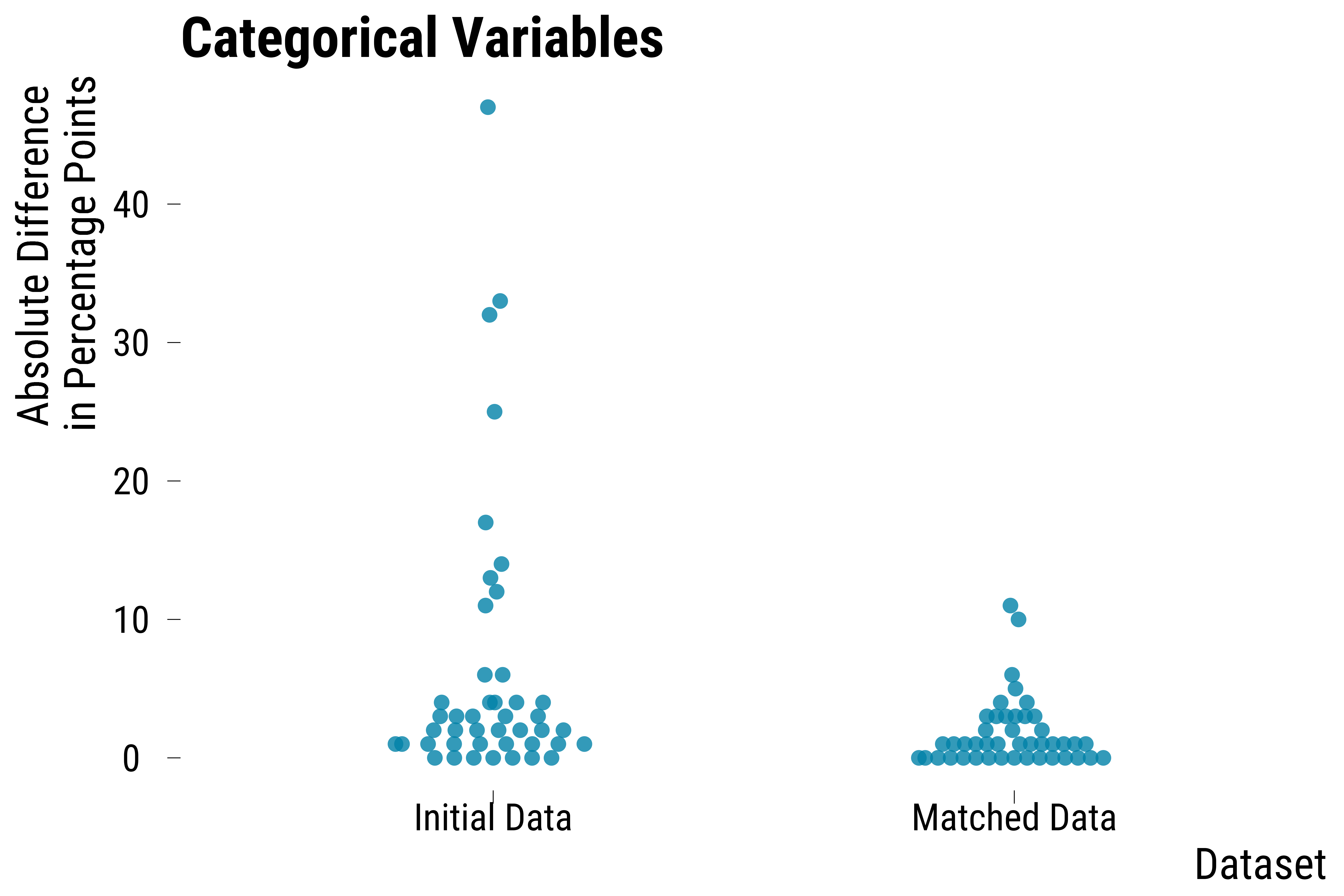
Categorical Covariates
Please show me the code!
# we select the dataset indicator and the standardized difference
data_calendar <- data_calendar %>%
ungroup() %>%
select(dataset, abs_difference)
data_weather_categorical <- data_weather_categorical %>%
ungroup() %>%
select(dataset, abs_difference)
data_categorical_love <-
bind_rows(data_calendar, data_weather_categorical)
# create the graph
graph_boxplot_categorical_balance_improvement <-
ggplot(data_categorical_love, aes(x = dataset, y = abs_difference)) +
ggbeeswarm::geom_quasirandom(
shape = 16,
size = 2,
width = 0.2,
color = my_blue,
alpha = 0.8
) +
scale_y_continuous(breaks = scales::pretty_breaks(n = 5)) +
xlab("Dataset") +
ylab("Absolute Difference \nin Percentage Points") +
ggtitle("Categorical Variables") +
theme_tufte() +
theme(axis.text.x = element_text(margin = ggplot2::margin(t = 0, unit = "cm")))
# print the graph
graph_boxplot_categorical_balance_improvement
Please show me the code!
# save the graph
ggsave(
graph_boxplot_categorical_balance_improvement,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_boxplot_categorical_balance_improvement.pdf"
),
width = 12,
height = 8,
units = "cm",
device = cairo_pdf
)
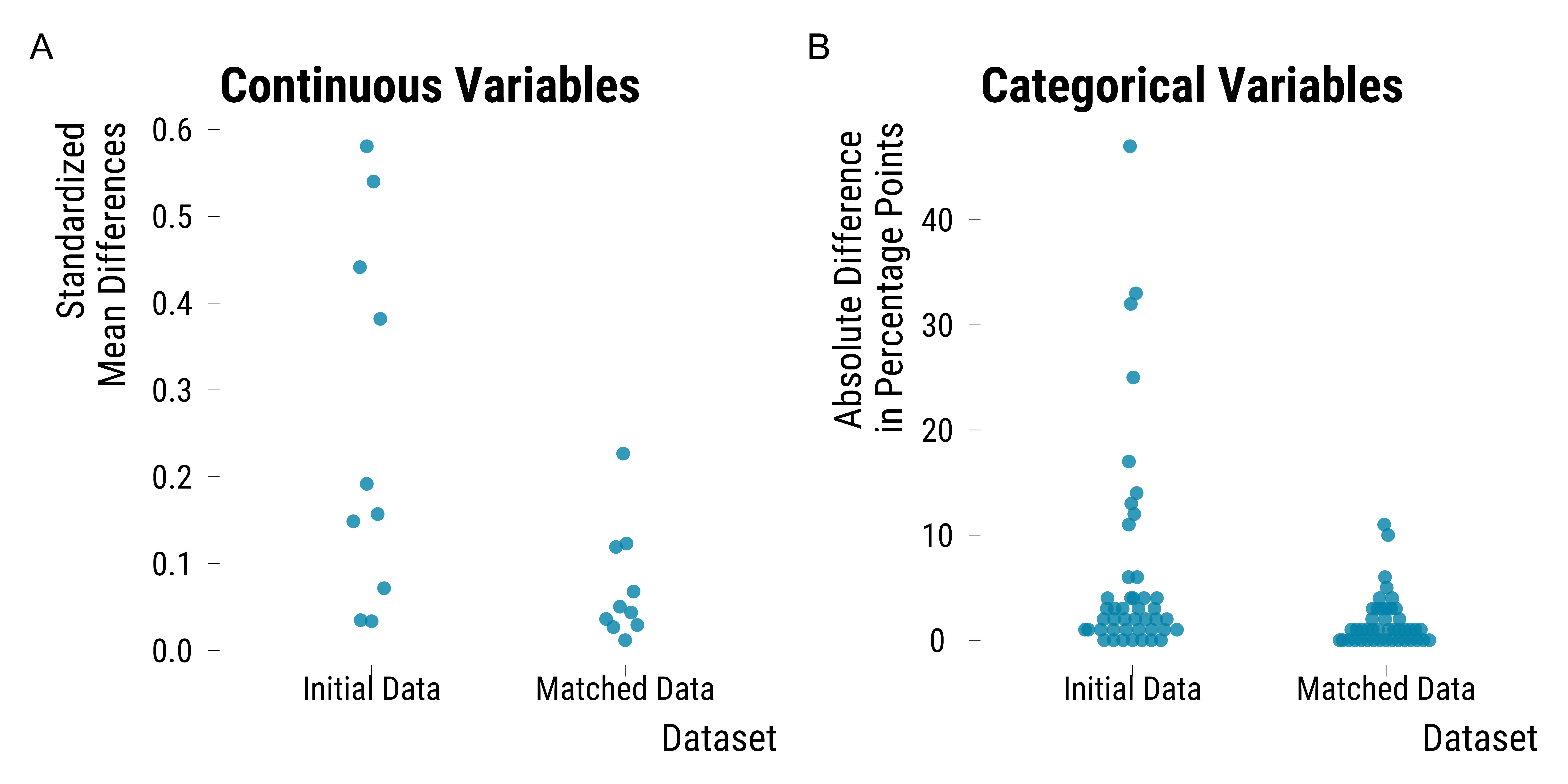
Combining Plots
We combine the two previous plots:
Please show me the code!
# combine the two previous plots
graph_overall_balance <-
graph_boxplot_continuous_balance_improvement + graph_boxplot_categorical_balance_improvement +
plot_annotation(tag_levels = 'A')
# display graph
graph_overall_balance
Please show me the code!
# save the graph
ggsave(
graph_overall_balance,
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_overall_balance.pdf"
),
width = 20,
height = 10,
units = "cm",
device = cairo_pdf
)
And we compute the overall figures for imbalance before and after matching:
Please show me the code!
# compute average imbalance before and after matching
data_categorical_love <- data_categorical_love %>%
mutate(Type = "Categorical (Difference in Percentage Points)") %>%
rename(standardized_difference = abs_difference)
data_continuous_love %>%
mutate(Type = "Continuous (Standardized Difference)") %>%
bind_rows(data_categorical_love) %>%
group_by(Type, dataset) %>%
summarise("Mean Imbalance" = round(mean(standardized_difference), 2)) %>%
rename(Dataset = dataset) %>%
knitr::kable(., align = c("l", "l", "c"))
| Type | Dataset | Mean Imbalance |
|---|---|---|
| Categorical (Difference in Percentage Points) | Initial Data | 6.23 |
| Categorical (Difference in Percentage Points) | Matched Data | 1.75 |
| Continuous (Standardized Difference) | Initial Data | 0.26 |
| Continuous (Standardized Difference) | Matched Data | 0.07 |
Randomization Check
Finally, we carry out a randomization check to statistically test whether the overall balance has increased after matching. As proposed by Gerber and Green (2012), we use as a balance measure the F-statistic from a regression where we regress the treatment indicator on all covariates.
We first compute the observed F-statistic for the initial data:
# compute f-statistic for initial data
fstat_initial_data <- data_matching %>%
lm(
is_treated ~ temperature_average + I(temperature_average ^ 2) +
temperature_average_lag_1 + I(temperature_average_lag_1 ^ 2) +
rainfall_duration + rainfall_duration_lag_1 +
humidity_average + humidity_average_lag_1 +
wind_speed + wind_speed_lag_1 +
weekday + holidays_dummy +
bank_day_dummy + month * as.factor(year),
data = .
) %>%
broom::glance() %>%
pull(statistic)
The F-statistic is equal to 8 for the initial data. We then freely permute 1000 times the treatment indicator to build the null distribution of the F-statistic:
Please show me the code!
# function to permute freely the treatment indicator
f_stat_free_permutation <- function(data) {
data %>%
mutate(is_treated = dqrng::dqsample(is_treated)) %>%
lm(
is_treated ~ temperature_average + I(temperature_average ^ 2) +
temperature_average_lag_1 + I(temperature_average_lag_1 ^ 2) +
rainfall_duration + rainfall_duration_lag_1 +
humidity_average + humidity_average_lag_1 +
wind_speed + wind_speed_lag_1 +
weekday + holidays_dummy +
bank_day_dummy + month * as.factor(year),
data = .
) %>%
broom::glance() %>%
pull(statistic)
}
# run 1000 permutations
data_f_stat_free_permutation <- tibble(sim_id = 1:1000) %>%
crossing(data_matching) %>%
group_by(sim_id) %>%
nest() %>%
mutate(f_stat = map(data, ~ f_stat_free_permutation(.))) %>%
select(-data) %>%
unnest(f_stat)
# save results
saveRDS(data_f_stat_free_permutation, here::here("inputs", "1.data", "5.matched_data", "data_f_stat_free_permutation.rds"))
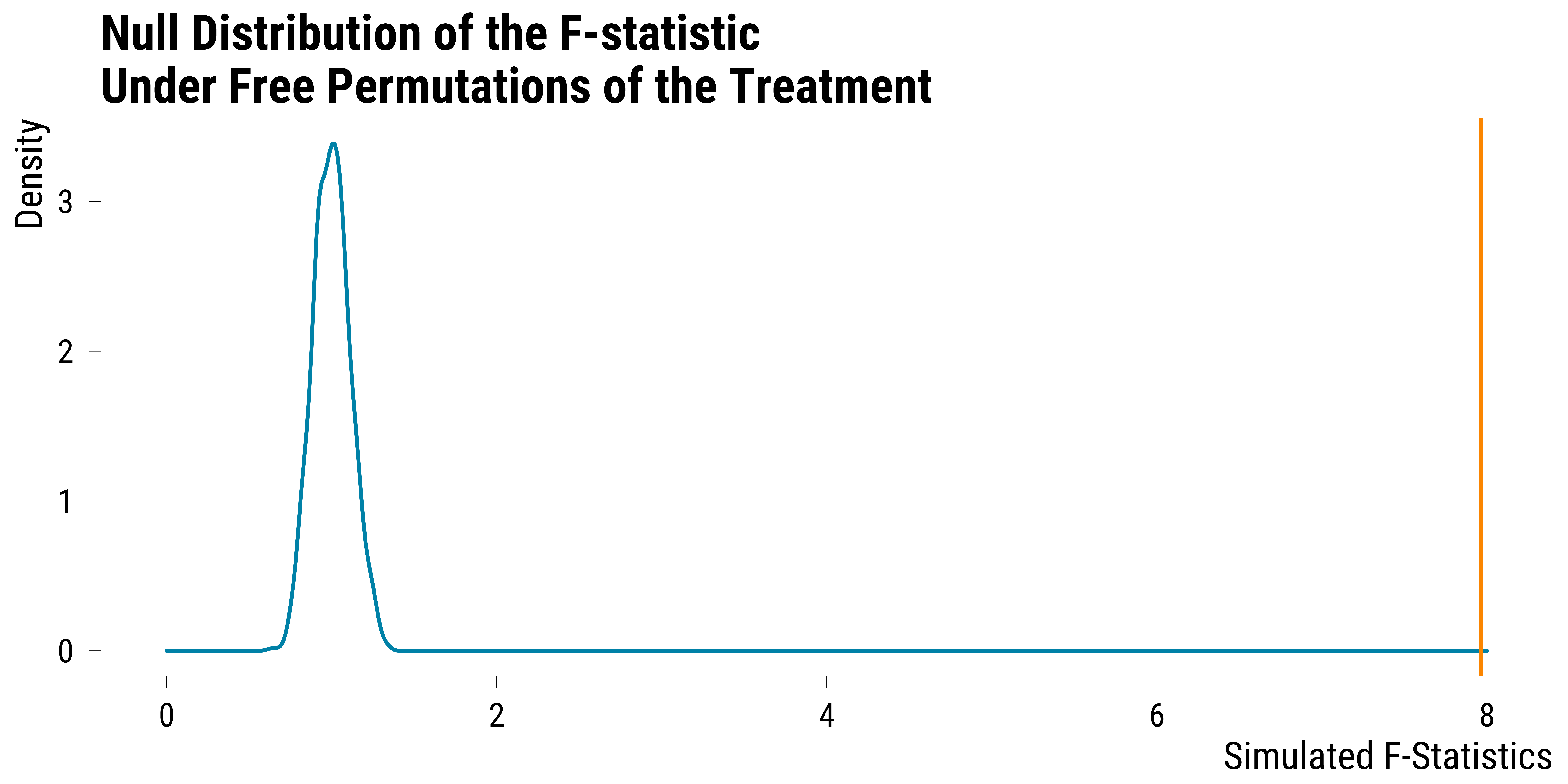
We plot the null distribution of the F-statistic:
Please show me the code!
# open results
data_f_stat_free_permutation <- readRDS(here::here("inputs", "1.data", "5.matched_data", "data_f_stat_free_permutation.rds"))
# make the graph
graph_f_stat_free_permutation <- data_f_stat_free_permutation %>%
ggplot(., aes(x = f_stat)) +
geom_density(colour = my_blue) +
geom_vline(xintercept = fstat_initial_data, colour = my_orange) +
xlim(0, 8) +
labs(x = "Simulated F-Statistics", y = "Density", title = "Null Distribution of the F-statistic\nUnder Free Permutations of the Treatment") +
theme_tufte()
# display the graph
graph_f_stat_free_permutation
Please show me the code!
# save the graph
ggsave(
graph_f_stat_free_permutation + labs(title = NULL),
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_balance_check_initial_data_1.pdf"
),
width = 15,
height = 10,
units = "cm",
device = cairo_pdf
)
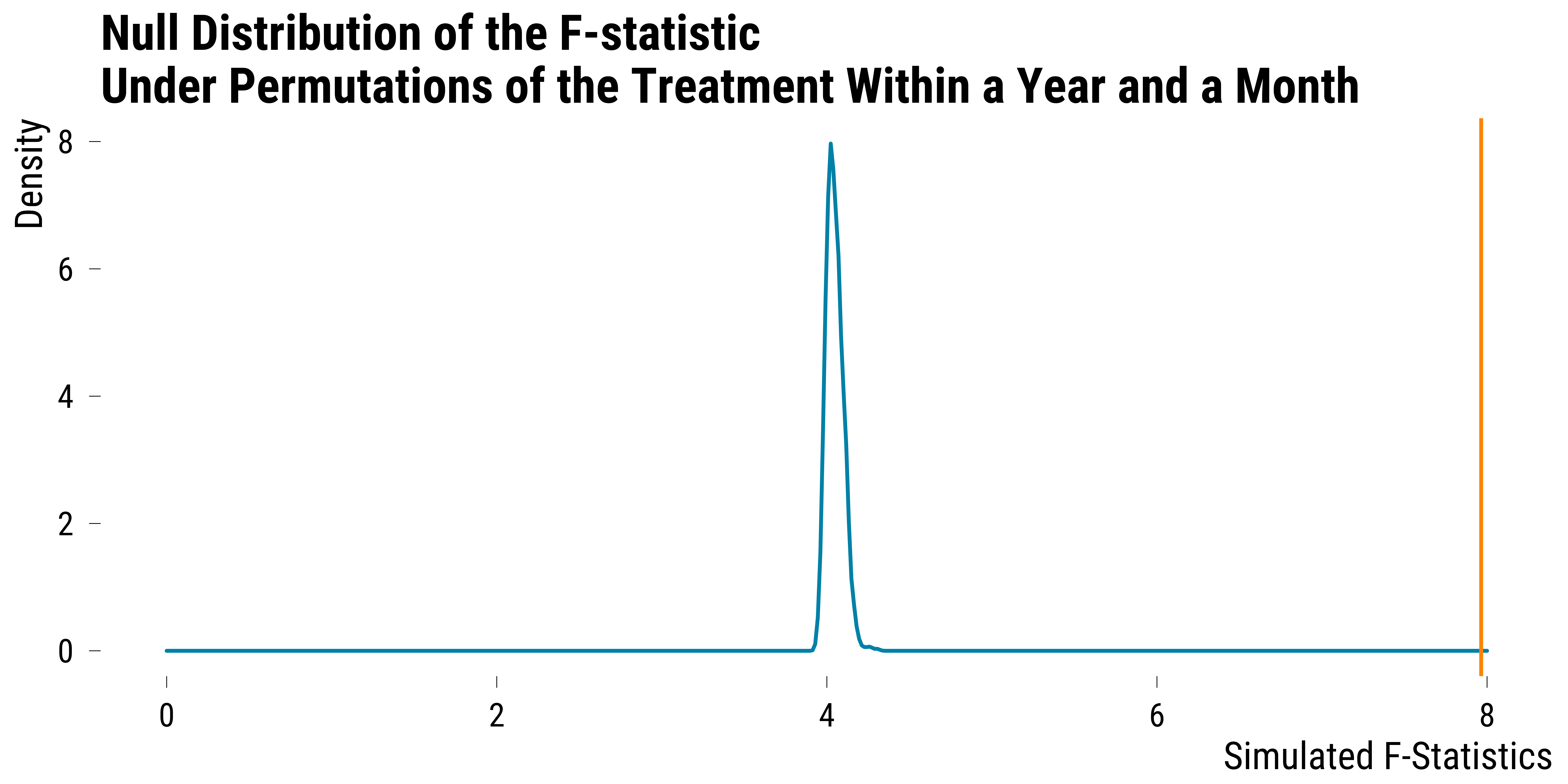
We can clearly reject the hypothesis that the no observed covariates have any effect on the treatment assignment. However, we freely permuted the treatment indicator, which removed the temporal structure of the data. To keep this temporal structure, we therefore carry out another balancing test where we permute the treatment indicator within a year and a month:
Please show me the code!
# function to permute by block the treatment indicator
f_stat_block_permutation <- function(data) {
data %>%
group_by(year, month) %>%
mutate(is_treated = dqrng::dqsample(is_treated)) %>%
lm(
is_treated ~ temperature_average + I(temperature_average ^ 2) +
temperature_average_lag_1 + I(temperature_average_lag_1 ^ 2) +
rainfall_duration + rainfall_duration_lag_1 +
humidity_average + humidity_average_lag_1 +
wind_speed + wind_speed_lag_1 +
weekday + holidays_dummy +
bank_day_dummy + month * as.factor(year),
data = .
) %>%
broom::glance() %>%
pull(statistic)
}
# run 1000 permutations
data_f_stat_block_permutation <- tibble(sim_id = 1:1000) %>%
crossing(data_matching) %>%
group_by(sim_id) %>%
nest() %>%
mutate(f_stat = map(data, ~ f_stat_block_permutation(.))) %>%
select(-data) %>%
unnest(f_stat)
# save results
saveRDS(data_f_stat_block_permutation, here::here("inputs", "1.data", "5.matched_data", "data_f_stat_block_permutation.rds"))
We plot the null distribution of the F-statistic:
Please show me the code!
# open results
data_f_stat_block_permutation <- readRDS(here::here("inputs", "1.data", "5.matched_data", "data_f_stat_block_permutation.rds"))
# make the graph
graph_f_stat_block_permutation <- data_f_stat_block_permutation %>%
ggplot(., aes(x = f_stat)) +
geom_density(colour = my_blue) +
geom_vline(xintercept = fstat_initial_data, colour = my_orange) +
xlim(0, 8) +
labs(x = "Simulated F-Statistics", y = "Density", title = "Null Distribution of the F-statistic\nUnder Permutations of the Treatment Within a Year and a Month") +
theme_tufte()
# display the graph
graph_f_stat_block_permutation
Please show me the code!
# save the graph
ggsave(
graph_f_stat_block_permutation + labs(title = NULL),
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_balance_check_initial_data_2.pdf"
),
width = 15,
height = 10,
units = "cm",
device = cairo_pdf
)
Again, we can reject the null hypothesis that no observed covariates have any effect on the treatment assignment.
We then compute the balance check for the matched data. We permute the treatment indicator within each pair as our matching procedure approximates a pairwise randomized experiment.
We compute the observed F-statistic for the matched data:
# compute f-statistic for matched data
fstat_matched_data <- data_matched %>%
lm(
is_treated ~ temperature_average + I(temperature_average ^ 2) +
temperature_average_lag_1 + I(temperature_average_lag_1 ^ 2) +
rainfall_duration + rainfall_duration_lag_1 +
humidity_average + humidity_average_lag_1 +
wind_speed + wind_speed_lag_1 +
weekday + holidays_dummy +
bank_day_dummy + month + as.factor(year),
data = .
) %>%
broom::glance() %>%
pull(statistic)
The F-statistic is equal to 0.5 for the matched data. We then implement our balance test:
Please show me the code!
# function to permute by pair the treatment indicator
f_stat_pair_permutation <- function(data) {
data %>%
group_by(pair_number) %>%
mutate(is_treated = dqrng::dqsample(is_treated)) %>%
lm(
is_treated ~ temperature_average + I(temperature_average ^ 2) +
temperature_average_lag_1 + I(temperature_average_lag_1 ^ 2) +
rainfall_duration + rainfall_duration_lag_1 +
humidity_average + humidity_average_lag_1 +
wind_speed + wind_speed_lag_1 +
weekday + holidays_dummy +
bank_day_dummy + month + as.factor(year),
data = .
) %>%
broom::glance() %>%
pull(statistic)
}
# run 1000 permutations
data_f_stat_matched_permutation <- tibble(sim_id = 1:1000) %>%
crossing(data_matched) %>%
group_by(sim_id) %>%
nest() %>%
mutate(f_stat = map(data, ~ f_stat_pair_permutation(.))) %>%
select(-data) %>%
unnest(f_stat)
# save results
saveRDS(data_f_stat_matched_permutation, here::here("inputs", "1.data", "5.matched_data", "data_f_stat_matched_permutation.rds"))
We plot the null distribution of the F-statistic:
Please show me the code!
# open results
data_f_stat_matched_permutation <- readRDS(here::here("inputs", "1.data", "5.matched_data", "data_f_stat_matched_permutation.rds"))
# make the graph
graph_f_stat_matched_permutation <- data_f_stat_matched_permutation %>%
ggplot(., aes(x = f_stat)) +
geom_density(colour = my_blue) +
geom_vline(xintercept = fstat_matched_data, colour = my_orange) +
xlim(0, 1.5) +
labs(x = "Simulated F-Statistics", y = "Density", title = "Null Distribution of the F-statistic\nUnder Permutations of the Treatment Within Pairs") +
theme_tufte()
# display graph
graph_f_stat_matched_permutation

Please show me the code!
# save the graph
ggsave(
graph_f_stat_matched_permutation + labs(title = NULL),
filename = here::here(
"inputs", "3.outputs",
"2.matching_analysis",
"graph_balance_check_matched_data.pdf"
),
width = 15,
height = 10,
units = "cm",
device = cairo_pdf
)
We can clearly see that the value of our observed F-statistic is likely to be obtained under the null hypothesis. Matching seems to have improve the covariates balance.